Variational Inference with Node Embeddings (VINE)
Siepel, Hassett, Staklinski — bioRxiv 2025
McCrone Lab Meeting
2025-01-08
Variational Inference
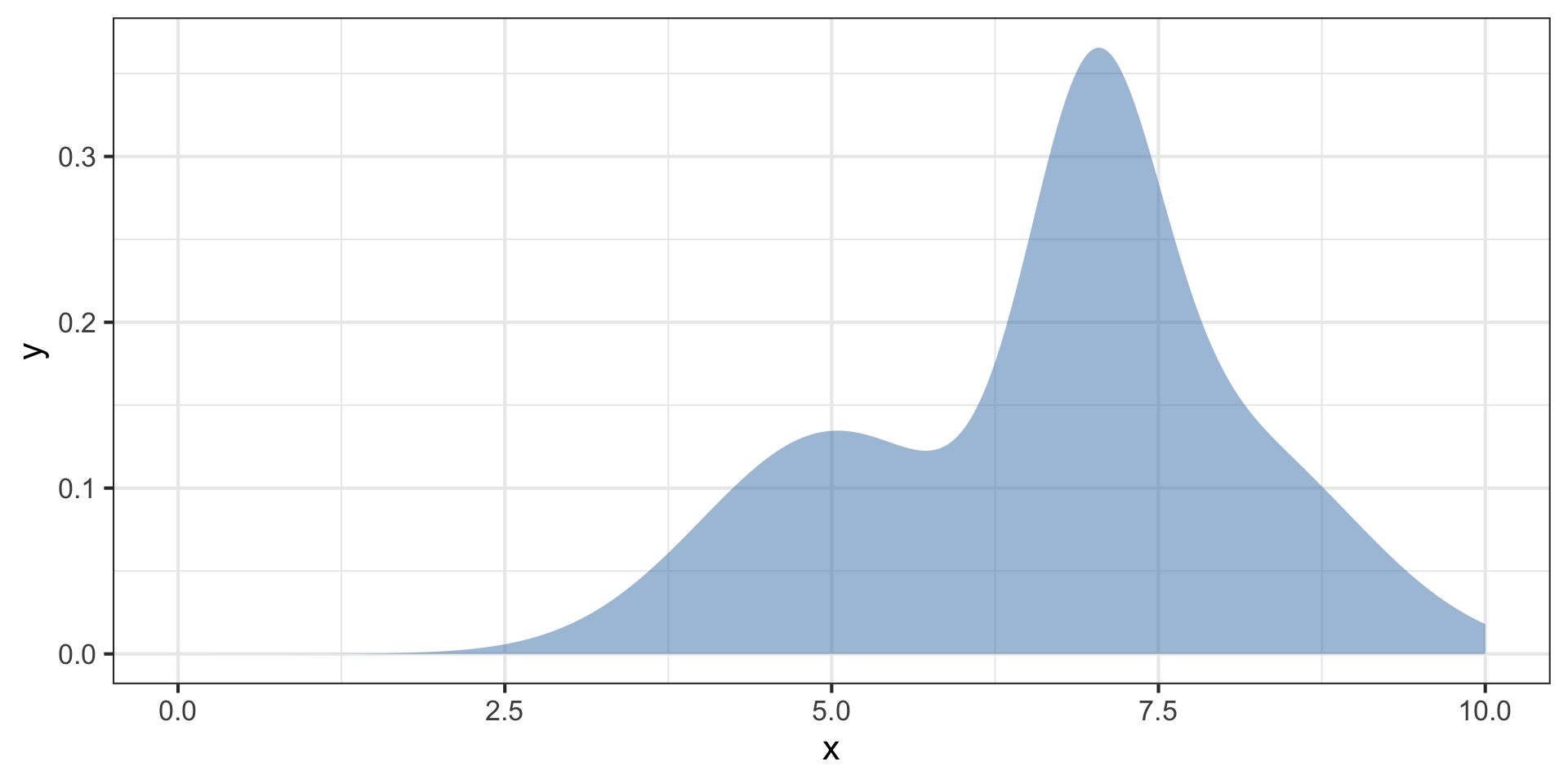
Posterior distributions are hard to characterize
We can use another function \(q(z)\) to approximate the posterior \(p(x,z)\).
Posterior distributions are hard to characterize
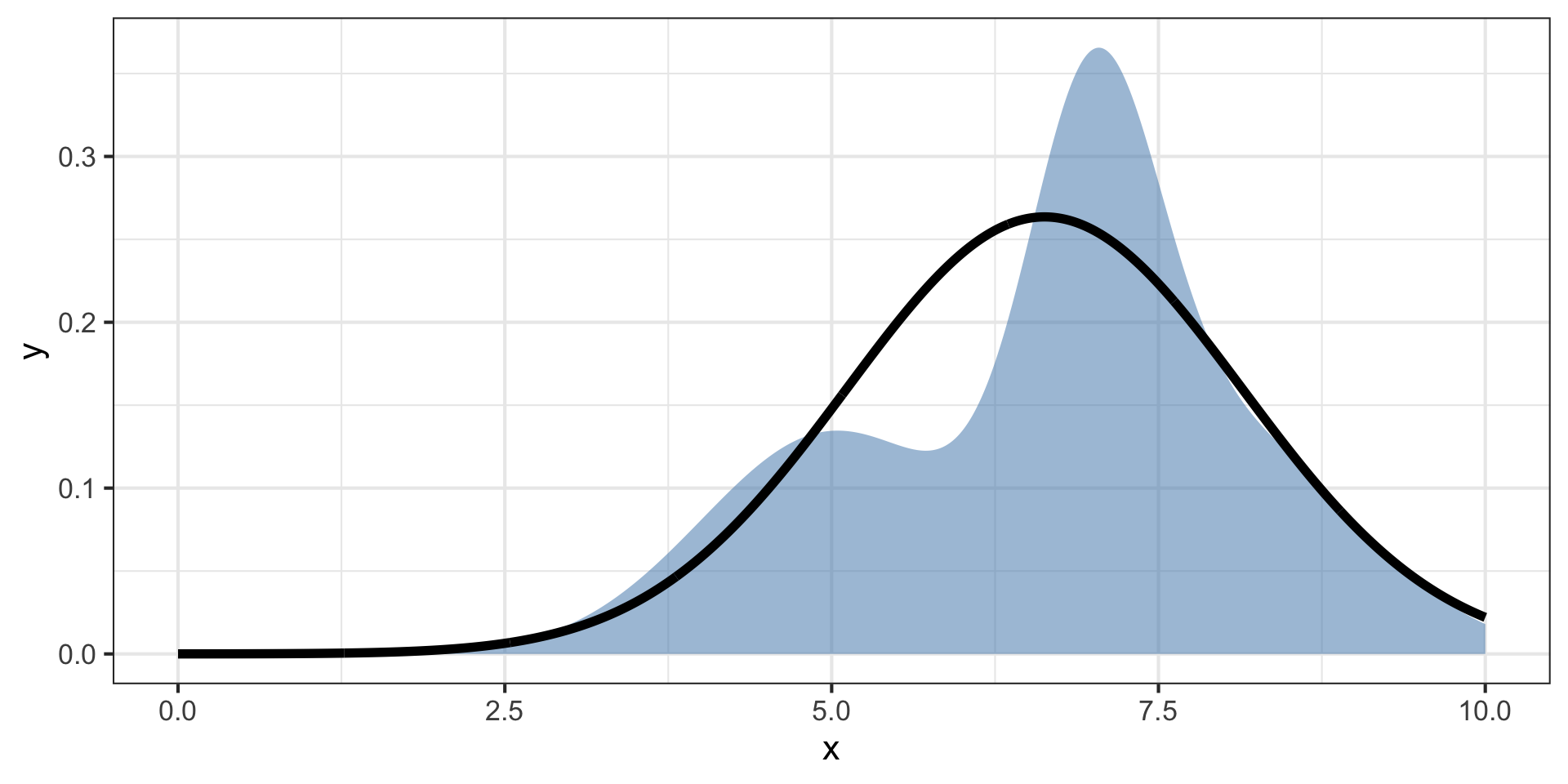
We can use another function \(q(z)\) to approximate the posterior \(p(x,z)\).
This is variational inference (VI)
Posterior distributions are hard to characterize
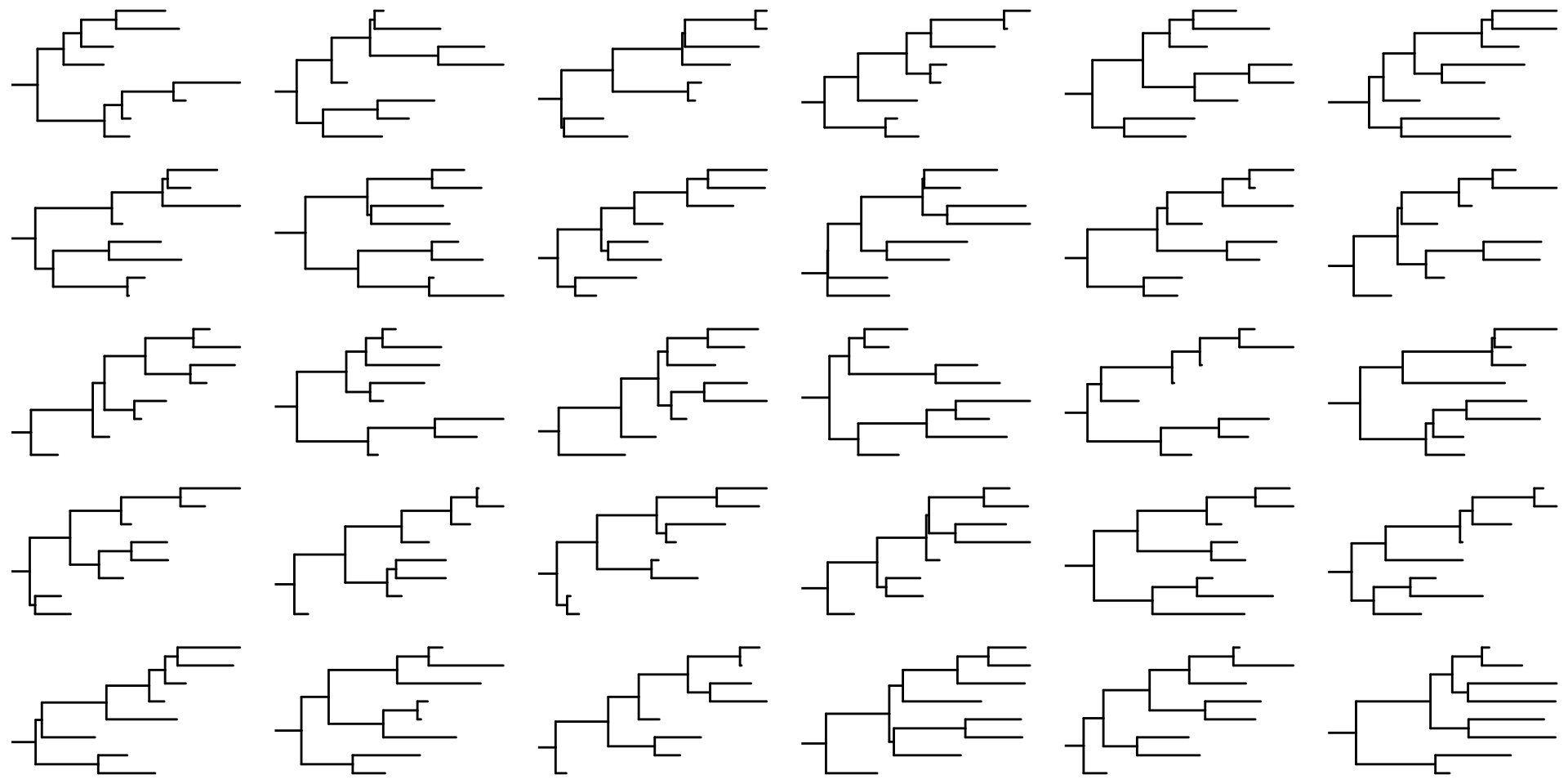
What if we could approximate the distribution of topologies and branch lengths with another distribution?
The central problem for VI on trees
Approximate the Bayesian posterior distribution of trees given a set of observed genotypes \(\mathbf{X}\) using a variational distribution \(q(\tau, \mathbf{b}; \theta)\)
\(\tau\) – topology of a tree
\(\mathbf{b}\) – vector of branch lengths
\([\)\(b_1\), \(b_2\), \(b_3\), \(b_4\)\(]\)
\(\theta\) – free parameters of the variational distribution
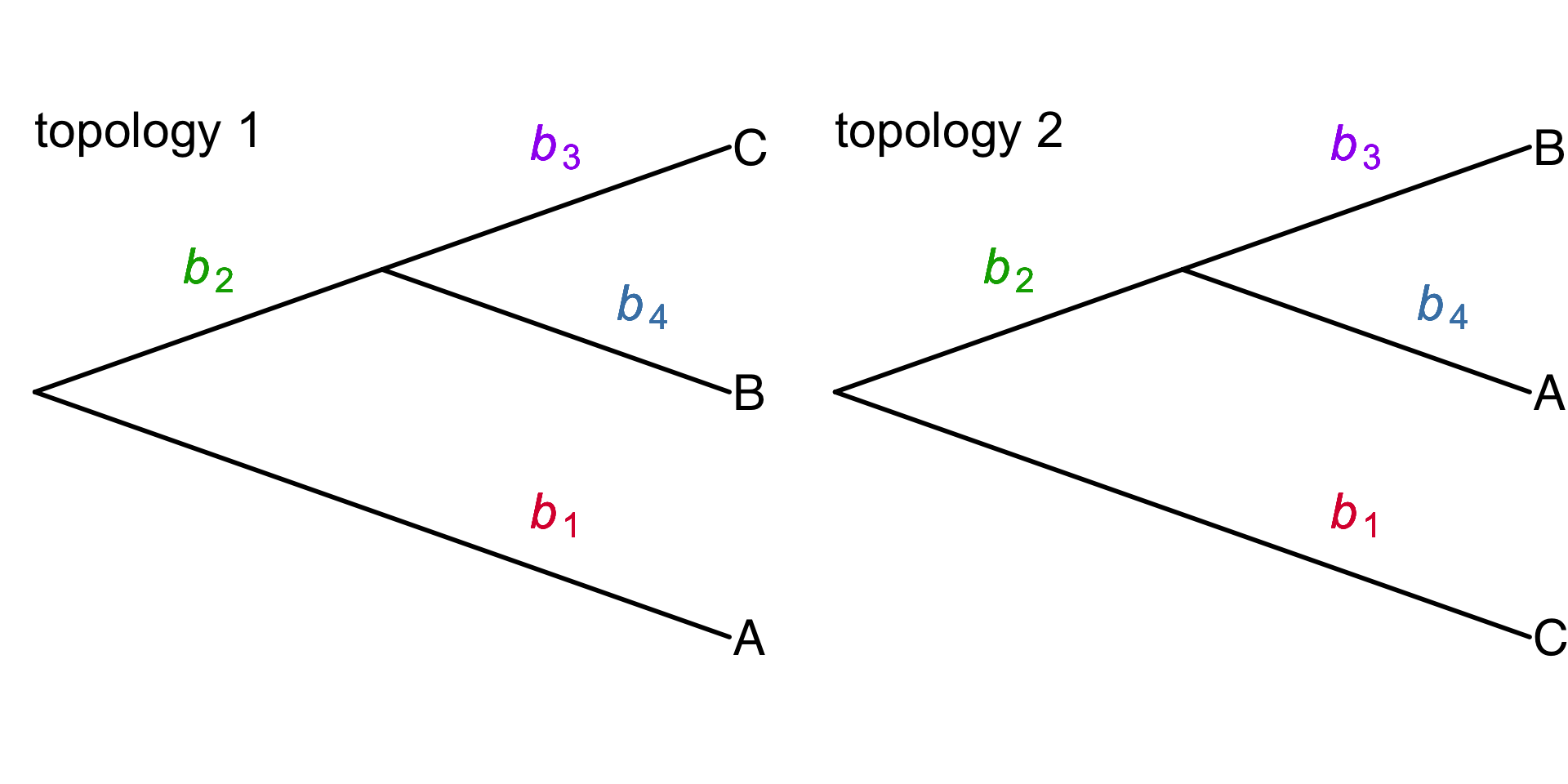
Typically, \(q\) is fitted to data by adjusting \(\theta\) to minimize the KL divergence from the true posterior distribution \(p(\tau,\mathbf{b} \mid X)\)
VI has been applied in phylogenetics before
- Variational Bayesian Phylogenetic Inference (VBPI) Cheng Zhang & Erick Matsen 2018
- VaiPhy Koptagel et al. 2022
- VBPI-Mixtures Kviman et al. 2023
- Dodonaphy Macaulay & Fourment 2024
- GeoPhy Mimori & Hamada 2023
Mimori & Hamada’s method
3 essential components
- Tips of the tree are embedded in a continuous space of \(d\) dimensions, with an induced a pairwise distance matrix \(\mathbf{D}\)
- Neighbor joining to convert \(\mathbf{D}\) to a tree with branch lengths \((\tau, \mathbf{b})\)
- Optimize the free parameters \(\theta\) of a simple variational distribution (multivariate normal) for the embedded points using stochastic gradient ascent.
VINE modifies Mimori & Hamada’s method in several ways
VINE – Variational Inference using Node Embeddings
- Tree topologies \(\tau\) an branch lengths \(\mathbf{b}\) are encoded in one continuous embedding
- Uses a standard (but accelerated) neighbor-joining (NJ) algorithm rather than a strictly continuous relaxation
- New algorithm for efficient backpropogation of gradients through the NJ algorithm
- Show that Taylor approximation of the objective function for VI results in nearly identical results as MCMC in a fraction of the time
- Introduce normalizing flows to accommodate nonlinearities in the approximate posterior distribution and richer parameterizations of the variational covariance matrix.
VINE steps
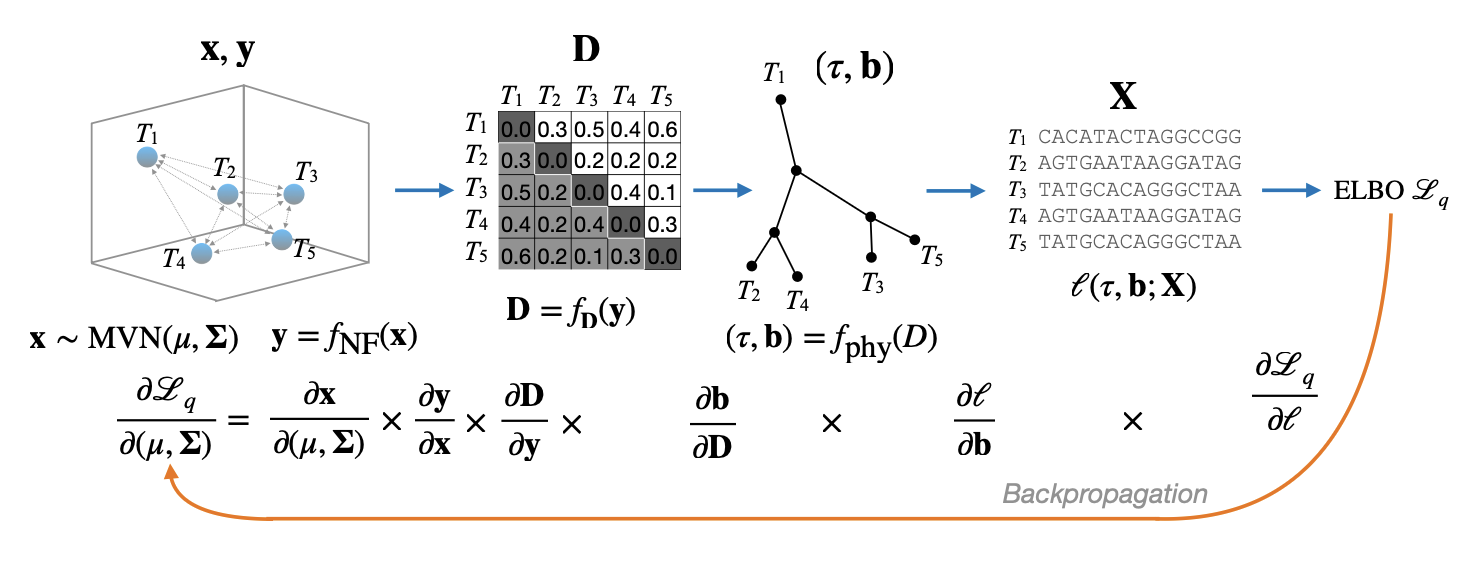
- Embedding – parameterized using a multivariate normal (\(\mathbb{R}^5\))
- Distance matrix is induced from the embedding
- Converted to a tree using neighbor joining
- Compute the likelihood of the branch lengths and topology given the sequences
- MCMC or Taylor Approximation of the ELBO (Evidence lower bound)
Gradients in neighbor joining?
Gradients in NJ
Observations
- NJ is piecewise smooth and differentiable almost everywhere
- small changes in the distance matrix \(D\) usually lead to smooth changes in branch lengths \(\mathbf{b}\)
- except when changes in \(D\) alter the sequence of neighbors leading to topology changes!
Neighbor joining has two steps
- sequence of selections of neighbors to be joined (A -> B) -> C
- mapping the distance matrix \(D\) to the branch length \(\mathbf{b}\) conditioned on these selections
NJ algorithm updates the matrices \(\mathbf{Q}\) and \(\mathbf{D}\)
If we condition on \(\mathbf{Q}\) (neighbor order), then only the distance matrix \(\mathbf{D}\) will contribute to the gradient.
They rely on stochastic gradient ascent to update the topologies as points migrate.
How do you get the gradient
NJ step 1: distance matrix update
After adding a new internal node, increase the size of the matrix by 1 and update distances.
Smooth, differentiable, and linear!
\(d^{(t+1)} = A^{(t)} d^{(t)}\)
NJ step 2: branch length update
I don’t want to TeX it out… but its also linear
\(b^{(t)} = C^{(t)} d^{(t)}\)